[1] 8R Functions & Regressions
A functional approach to programming
March 18, 2026
Functions
Functions
Functions…
take input(s)
do something
return an ouput
R is a Functional Language
R, at its heart, is a functional programming (FP) language. This means that it provides many tools for the creation and manipulation of functions. In particular, R has what’s known as first class functions. You can do anything with functions that you can do with vectors: you can assign them to variables, store them in lists, pass them as arguments to other functions, create them inside functions, and even return them as the result of a function. Hadley Wickham, Advanced R
Why use functions?
Functions make your code:
more flexible
less repetitive
more readable (potentially)
A Basic Function
Here is a basic, if not exciting, function:
- Take inputs
- Do something
- Return output
Syntax
Functions are always declared using the function function(),
followed by curly braces { } which demark what the function actually does.
Most functions will end with a return() call, though this is not strictly required.
“Degenerate” functions
Which is just to show that technically inputs and outputs are not necessary.
Variable Scope
Functions have their own “environment” for variables.
[1] "Outside function"Even if we call our function, the value for cubed is not overwritten in our R session environment.
Breaking Function Scope
<<- breaks the scope of a function and affects vaiables outside
[1] 27[1] 27But this is a really bad idea.
You should only allow functions to affect your session by returning values.
Otherwise, it is very confusing to tell what is changing a variable.
Limiting Function Scope
Technically, you can use a variable defined outside a function inside a function.
But this is also a bad idea.
Functions are meant to be flexible and portable
By relying on a session variable we’ve made this function dependent on the current setting.
Adding Additional Arguments
Instead, we should pass beta as an additional argument for our function.
Setting a Default Argument
When declaring a function, you can set a default value for an argument.
That default value will be used unless you specify a new value to overwrite it.
Ordering of Arguments
Functions assume unlabelled arguments are given in the order they were declared.
i.e. the first argument is our consumption value, and the second is our beta.
You can always be explicit about function arguments as well.
CRRA Utility
Let’s declare a CRRA utility function,
\[ U(C) = \beta \frac{C^{1-\gamma}}{1-\gamma} \]
In our code, this looks like
CRRA Utility
Let’s declare a CRRA utility function,
\[ U(C) = \beta \frac{C^{1-\gamma}}{1-\gamma} \]
What happens if we run when gamma = 1?
If you paid attention in Macro, then you know that CRRA utility with \(\gamma \rightarrow 1\) converges to log utility.
If-Else Statements within Functions
We can implement this with an if-else statement within our function.
Now our function works in both cases:
Printing out a function
We can always inspect the actual code for a function by calling it without the parentheses ().
Printing out a function
This works for functions from other packages too.
function (x, n = 1L, default = NULL, order_by = NULL, ...)
{
if (inherits(x, "ts")) {
abort("`x` must be a vector, not a <ts>, do you want `stats::lag()`?")
}
check_dots_empty0(...)
check_number_whole(n)
if (n < 0L) {
abort("`n` must be positive.")
}
shift(x, n = n, default = default, order_by = order_by)
}
<bytecode: 0x10a27c250>
<environment: namespace:dplyr>Anonymous Functions
Anonymous functions are functions that do not have a name.

Anonymous Functions
Anonymous functions are functions that do not have a name.
This means they are not stored as a variable that you can use over again, but exist only for a moment.
For functions that only take one line, you can drop the curly brackets.
When would we use this?
purrr::map() family
Map Family of Functions
We saw briefly the *apply() family of base R functions (when we were looking at loops).
The map() family is the tidyverse equivalent and are nicer to use.
Map Function
The map() function always
- takes a vector (or list) for input
- calls a function on each element
- returns a list as the result
Using an Anonymous Function with map()
This is an example where we may want to declare an anonymous function.
Using an Anonymous Function with map()
And again, because our function is only one line, we could drop the curly braces.
Multiline Anonymous Functions
Conversely, you could have a multiline anonymous function.
[[1]]
mpg cyl disp
Min. :10.40 Min. :4.000 Min. : 71.1
1st Qu.:15.43 1st Qu.:4.000 1st Qu.:120.8
Median :19.20 Median :6.000 Median :196.3
Mean :20.09 Mean :6.188 Mean :230.7
3rd Qu.:22.80 3rd Qu.:8.000 3rd Qu.:326.0
Max. :33.90 Max. :8.000 Max. :472.0
[[2]]
Sepal.Length Sepal.Width Petal.Length
Min. :4.300 Min. :2.000 Min. :1.000
1st Qu.:5.100 1st Qu.:2.800 1st Qu.:1.600
Median :5.800 Median :3.000 Median :4.350
Mean :5.843 Mean :3.057 Mean :3.758
3rd Qu.:6.400 3rd Qu.:3.300 3rd Qu.:5.100
Max. :7.900 Max. :4.400 Max. :6.900 map Return Types
By default, map() will always return a list.
It does not know the datatype of the objects you are returning,
- and a list works with any data type and with a mix of data types.
If you do know your datatype, you could use…
map_lgl()map_int()map_dbl()map_chr()
- Which will all return a vector of that data type.
Example with map_dbl()
Using our prior map example,
Example with fredr package
We saw before that map() was convenient for working with fredr package.
This map function returns data.frames (tibbles) and then list_rbind combines them by rbind-ing them.
rbind()takes two data.frames and stacks them on top of each othercbind()takes two data.frames and stacks them beside one another
Regression Functions
Regressions
The workhorse of economics research is linear regression.
The simplest form of which is, \[ Y = \beta X + \epsilon \]
Given a set of values for \(Y\) and \(X\), we can estimate \(\hat{\beta}\).
Generating some example data
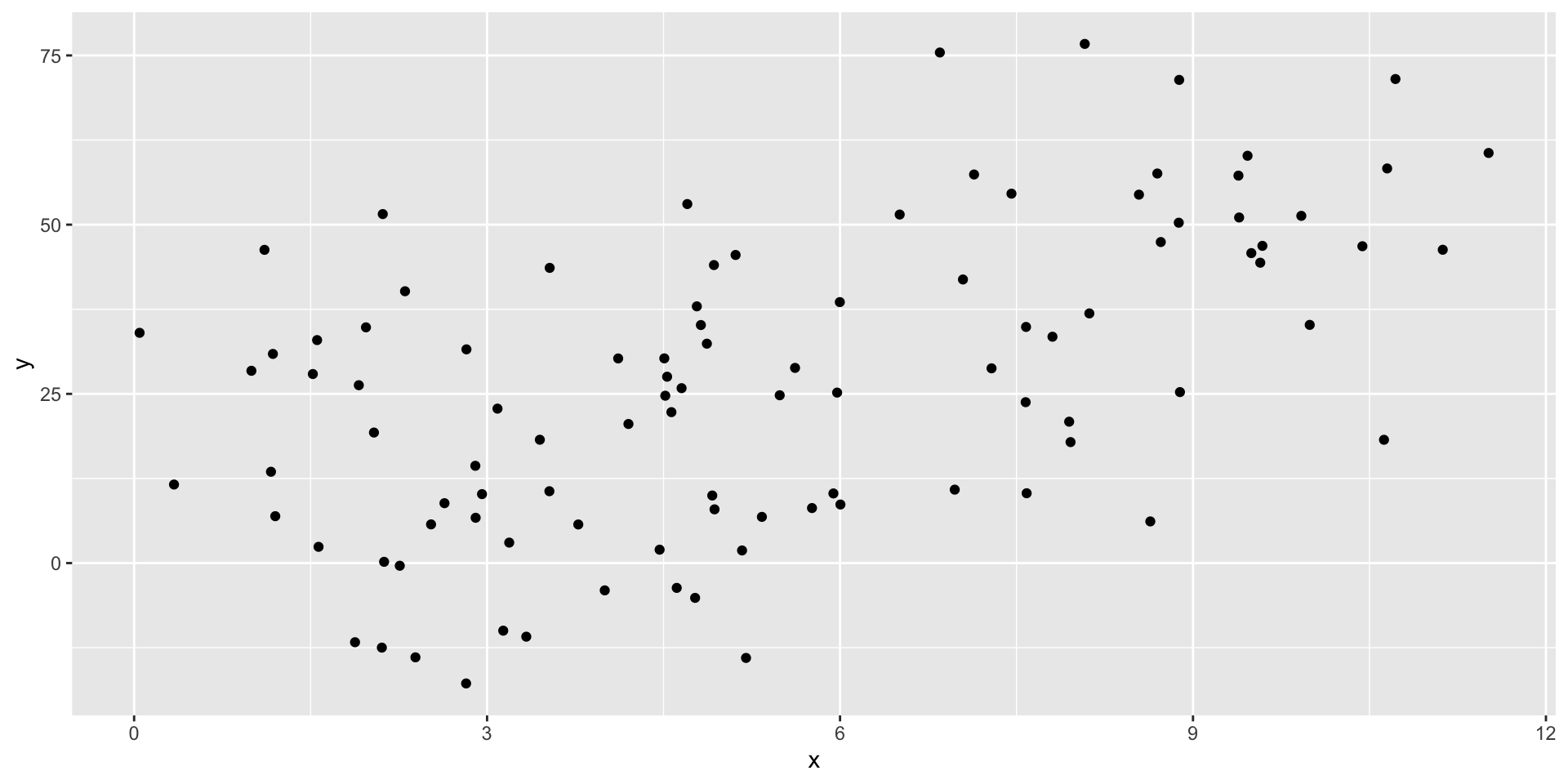
Estimating \(\beta\)
R has a built in function for linear models: lm().
summary() for detailed information
Call:
lm(formula = "y ~ x", data = tb)
Residuals:
Min 1Q Median 3Q Max
-39.971 -14.708 -0.024 13.895 42.721
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 4.633 3.939 1.176 0.242
x 4.100 0.643 6.377 5.97e-09 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 18.85 on 98 degrees of freedom
Multiple R-squared: 0.2932, Adjusted R-squared: 0.286
F-statistic: 40.66 on 1 and 98 DF, p-value: 5.973e-09Structure of lm() models
List of 12
$ coefficients : Named num [1:2] 4.63 4.1
..- attr(*, "names")= chr [1:2] "(Intercept)" "x"
$ residuals : Named num [1:100] 26.09 -19.69 6.29 5.99 22.93 ...
..- attr(*, "names")= chr [1:100] "1" "2" "3" "4" ...
$ effects : Named num [1:100] -266.88 -120.21 1.64 7.22 24.75 ...
..- attr(*, "names")= chr [1:100] "(Intercept)" "x" "" "" ...
$ rank : int 2
$ fitted.values: Named num [1:100] 14.1 26.5 13 45.3 48.6 ...
..- attr(*, "names")= chr [1:100] "1" "2" "3" "4" ...
$ assign : int [1:2] 0 1
$ qr :List of 5
..$ qr : num [1:100, 1:2] -10 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 ...
.. ..- attr(*, "dimnames")=List of 2
.. .. ..$ : chr [1:100] "1" "2" "3" "4" ...
.. .. ..$ : chr [1:2] "(Intercept)" "x"
.. ..- attr(*, "assign")= int [1:2] 0 1
..$ qraux: num [1:2] 1.1 1.01
..$ pivot: int [1:2] 1 2
..$ tol : num 1e-07
..$ rank : int 2
..- attr(*, "class")= chr "qr"
$ df.residual : int 98
$ xlevels : Named list()
$ call : language lm(formula = "y ~ x", data = tb)
$ terms :Classes 'terms', 'formula' language y ~ x
.. ..- attr(*, "variables")= language list(y, x)
.. ..- attr(*, "factors")= int [1:2, 1] 0 1
.. .. ..- attr(*, "dimnames")=List of 2
.. .. .. ..$ : chr [1:2] "y" "x"
.. .. .. ..$ : chr "x"
.. ..- attr(*, "term.labels")= chr "x"
.. ..- attr(*, "order")= int 1
.. ..- attr(*, "intercept")= int 1
.. ..- attr(*, "response")= int 1
.. ..- attr(*, ".Environment")=<environment: 0x13d181e98>
.. ..- attr(*, "predvars")= language list(y, x)
.. ..- attr(*, "dataClasses")= Named chr [1:2] "numeric" "numeric"
.. .. ..- attr(*, "names")= chr [1:2] "y" "x"
$ model :'data.frame': 100 obs. of 2 variables:
..$ y: num [1:100] 40.16 6.83 19.29 51.3 71.52 ...
..$ x: num [1:100] 2.3 5.34 2.04 9.92 10.72 ...
..- attr(*, "terms")=Classes 'terms', 'formula' language y ~ x
.. .. ..- attr(*, "variables")= language list(y, x)
.. .. ..- attr(*, "factors")= int [1:2, 1] 0 1
.. .. .. ..- attr(*, "dimnames")=List of 2
.. .. .. .. ..$ : chr [1:2] "y" "x"
.. .. .. .. ..$ : chr "x"
.. .. ..- attr(*, "term.labels")= chr "x"
.. .. ..- attr(*, "order")= int 1
.. .. ..- attr(*, "intercept")= int 1
.. .. ..- attr(*, "response")= int 1
.. .. ..- attr(*, ".Environment")=<environment: 0x13d181e98>
.. .. ..- attr(*, "predvars")= language list(y, x)
.. .. ..- attr(*, "dataClasses")= Named chr [1:2] "numeric" "numeric"
.. .. .. ..- attr(*, "names")= chr [1:2] "y" "x"
- attr(*, "class")= chr "lm"It’s just a big list!
You can access certain components (ex. m$residuals).
Summary also returns a list
List of 11
$ call : language lm(formula = "y ~ x", data = tb)
$ terms :Classes 'terms', 'formula' language y ~ x
.. ..- attr(*, "variables")= language list(y, x)
.. ..- attr(*, "factors")= int [1:2, 1] 0 1
.. .. ..- attr(*, "dimnames")=List of 2
.. .. .. ..$ : chr [1:2] "y" "x"
.. .. .. ..$ : chr "x"
.. ..- attr(*, "term.labels")= chr "x"
.. ..- attr(*, "order")= int 1
.. ..- attr(*, "intercept")= int 1
.. ..- attr(*, "response")= int 1
.. ..- attr(*, ".Environment")=<environment: 0x13d181e98>
.. ..- attr(*, "predvars")= language list(y, x)
.. ..- attr(*, "dataClasses")= Named chr [1:2] "numeric" "numeric"
.. .. ..- attr(*, "names")= chr [1:2] "y" "x"
$ residuals : Named num [1:100] 26.09 -19.69 6.29 5.99 22.93 ...
..- attr(*, "names")= chr [1:100] "1" "2" "3" "4" ...
$ coefficients : num [1:2, 1:4] 4.633 4.1 3.939 0.643 1.176 ...
..- attr(*, "dimnames")=List of 2
.. ..$ : chr [1:2] "(Intercept)" "x"
.. ..$ : chr [1:4] "Estimate" "Std. Error" "t value" "Pr(>|t|)"
$ aliased : Named logi [1:2] FALSE FALSE
..- attr(*, "names")= chr [1:2] "(Intercept)" "x"
$ sigma : num 18.9
$ df : int [1:3] 2 98 2
$ r.squared : num 0.293
$ adj.r.squared: num 0.286
$ fstatistic : Named num [1:3] 40.7 1 98
..- attr(*, "names")= chr [1:3] "value" "numdf" "dendf"
$ cov.unscaled : num [1:2, 1:2] 0.04366 -0.00626 -0.00626 0.00116
..- attr(*, "dimnames")=List of 2
.. ..$ : chr [1:2] "(Intercept)" "x"
.. ..$ : chr [1:2] "(Intercept)" "x"
- attr(*, "class")= chr "summary.lm"It can be useful to access specific components with summary(m)$coefficients.
Tidying model results
broom is a package that tries to “tidy” model results into data.frames.
Let’s see what it does for our little model:
# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 4.63 3.94 1.18 0.242
2 x 4.10 0.643 6.38 0.00000000597Much easier to work with!
broom is written to create standardized results from many different models from many different packages.
Subsets of data
First, we are going to add some random groups to our data.
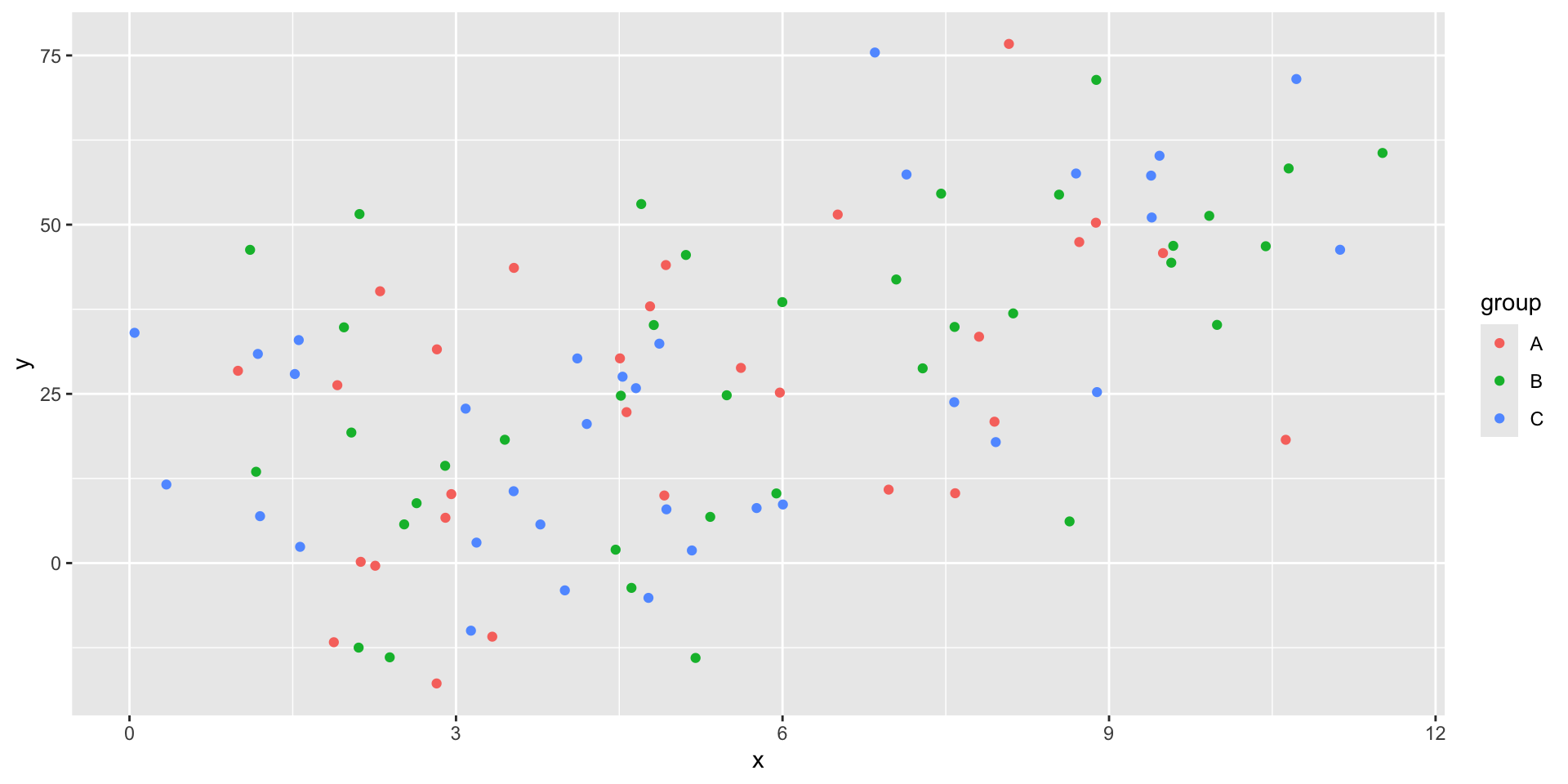
Nesting tibbles
What if we want to run multiple regressions, one on each group?
One option would be filter the data, then call lm three times.
Running lm on nested tibbles
Now we can use this nested format to run all three of our models at once.
# A tibble: 3 × 3
group data model
<chr> <list> <list>
1 A <tibble [29 × 2]> <lm>
2 B <tibble [37 × 2]> <lm>
3 C <tibble [34 × 2]> <lm> Now we have a column of lm models!
Tidying nested model results
Now we can tidy those models so we can contrast all of the results.
# A tibble: 3 × 4
group data model summary
<chr> <list> <list> <list>
1 A <tibble [29 × 2]> <lm> <tibble [2 × 5]>
2 B <tibble [37 × 2]> <lm> <tibble [2 × 5]>
3 C <tibble [34 × 2]> <lm> <tibble [2 × 5]>We have all of the model summaries now, but we need to unnest the tibble to easily compare them.
Unnesting a tibble
# A tibble: 6 × 6
group term estimate std.error statistic p.value
<chr> <chr> <dbl> <dbl> <dbl> <dbl>
1 A (Intercept) 5.42 7.90 0.686 0.499
2 B (Intercept) 5.33 6.92 0.770 0.447
3 C (Intercept) 3.88 6.36 0.609 0.547
4 A x 3.74 1.38 2.72 0.0113
5 B x 4.10 1.05 3.89 0.000432
6 C x 4.27 1.07 4.00 0.000354We can now easily compare our three models’ \(\beta\) estimate for \(x\).
Making a coefficient chart
stb |>
unnest(summary) |>
ggplot() +
geom_point(aes(
x = estimate,
y = group,
color = group
),
size = 3
) +
geom_segment(aes(
x = estimate - 1.96 * std.error,
xend = estimate + 1.96 * std.error,
y = group,
yend = group,
color = group
)) +
geom_vline(xintercept = 0) +
facet_wrap(vars(term), scales = "free")Making a coefficient chart
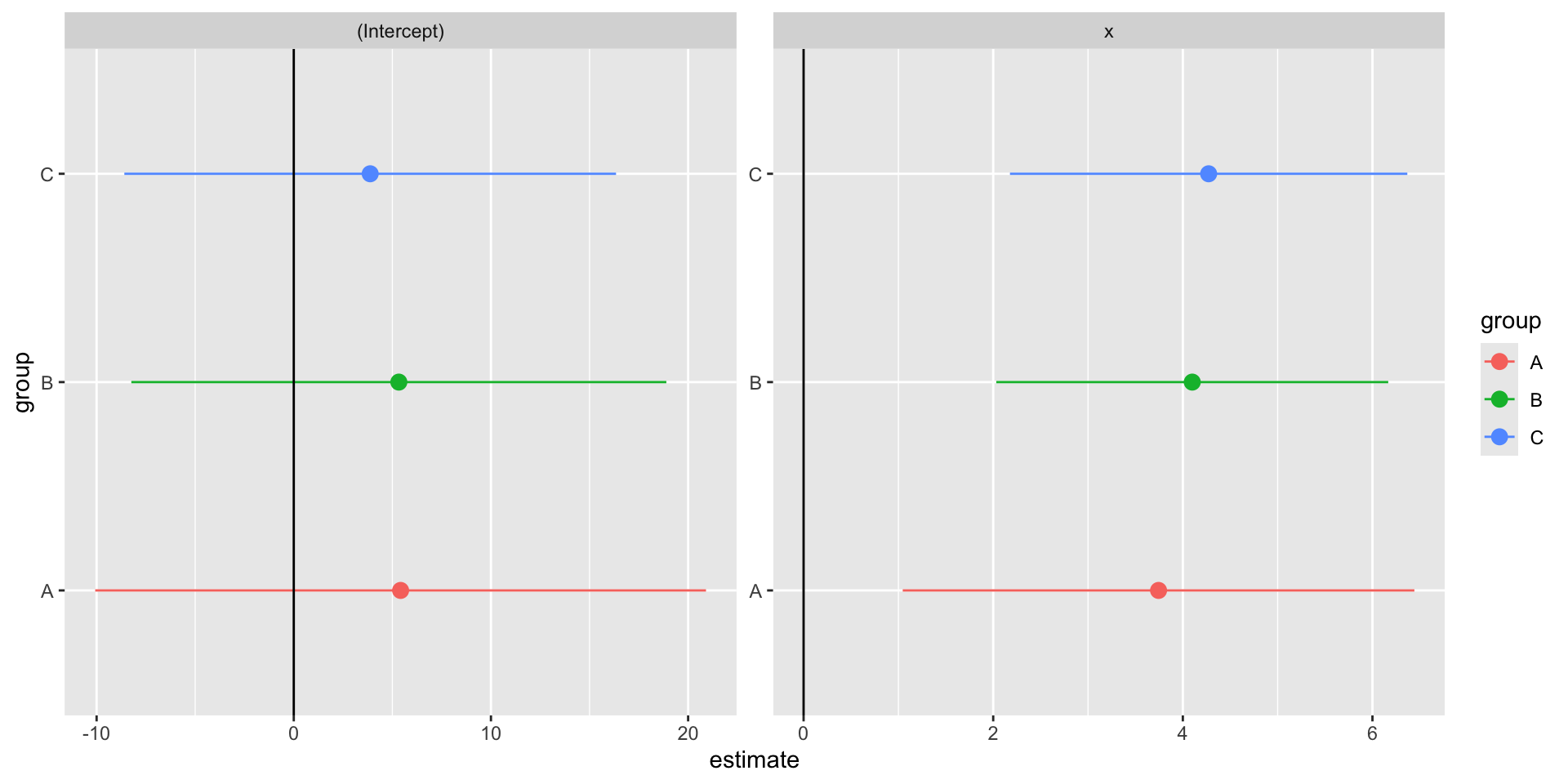
Nesting Summary
Pros
A clean way to estimate multiple models
Makes full use of
tidyversefunctionsEasy to add more models, different subsets
Cons
- Duplicates data for each row
- This can be very costly for large datasets
Robust Standard Errors
Robust Standard Errors
As you know from econometrics class, you shold pretty much always use robust standard errors when you report results.
There are a couple of packages which support this in R,
Robust SE with estimatr
The package is written to match the syntax of lm().
Same point estimates, slightly different standard errors.
# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 4.63 3.94 1.18 0.242
2 x 4.10 0.643 6.38 0.00000000597 term estimate std.error statistic p.value conf.low conf.high df
1 (Intercept) 4.633091 4.0673715 1.139087 2.574435e-01 -3.438476 12.70466 98
2 x 4.100106 0.6251976 6.558096 2.574586e-09 2.859422 5.34079 98
outcome
1 y
2 yChanging the SE type
estimatr defaults to using “HC2” robust standard errors.
There are options for “HC0”, “HC1”, and “HC3”.
I can’t really tell you the importance of the small differences between these robust options. You can read more about the math for each here.
Panel Regressions
Panel Regressions
Fixed effects
Clustering
Panel data
Panel data has at least two dimensions
- time
- groups
- countries, states, people
This is in contrast to time series (just a time dimension) and cross-sectional data.
Example Panel Data
We are going to use the package gapminder to get our example panel data.
# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <dbl> <dbl>
1 Afghanistan Asia 1952 28.8 8.43 0.779
2 Afghanistan Asia 1957 30.3 9.24 0.821
3 Afghanistan Asia 1962 32.0 10.3 0.853
4 Afghanistan Asia 1967 34.0 11.5 0.836
5 Afghanistan Asia 1972 36.1 13.1 0.740
6 Afghanistan Asia 1977 38.4 14.9 0.786
7 Afghanistan Asia 1982 39.9 12.9 0.978
8 Afghanistan Asia 1987 40.8 13.9 0.852
9 Afghanistan Asia 1992 41.7 16.3 0.649
10 Afghanistan Asia 1997 41.8 22.2 0.635
# ℹ 1,694 more rowsRunning OLS with fixest
Let’s predict life expectancy using GDP per capita.
OLS estimation, Dep. Var.: lifeExp
Observations: 1,704
Standard-errors: IID
Estimate Std. Error t value Pr(>|t|)
(Intercept) 53.955561 0.314995 171.2902 < 2.2e-16 ***
gdpPercap 0.764883 0.025790 29.6577 < 2.2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
RMSE: 10.5 Adj. R2: 0.340326We could easily have estimated this with
lm()orlm_robust()
Adding in fixed effects
Let’s add fixed effects for each continent.
OLS estimation, Dep. Var.: lifeExp
Observations: 1,704
Fixed-effects: continent: 5
Standard-errors: IID
Estimate Std. Error t value Pr(>|t|)
gdpPercap 0.44527 0.023498 18.9493 < 2.2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
RMSE: 8.37531 Adj. R2: 0.578106
Within R2: 0.174557Super easy to run!
Notice that it automatically clustered our standard errors to “continent” as well. What if we don’t want that? When Should You Adjust Standard Errors for Clustering (Abadie, Athey, Imbens, Woolridge 2017)
Setting iid standard errors
We can be explicit about standard errors using the “vcov=” argument.
OLS estimation, Dep. Var.: lifeExp
Observations: 1,704
Fixed-effects: continent: 5
Standard-errors: IID
Estimate Std. Error t value Pr(>|t|)
gdpPercap 0.44527 0.023498 18.9493 < 2.2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
RMSE: 8.37531 Adj. R2: 0.578106
Within R2: 0.174557Setting iid standard errors
We can also have clustered standard errors without including fixed effects.
OLS estimation, Dep. Var.: lifeExp
Observations: 1,704
Standard-errors: Clustered (continent)
Estimate Std. Error t value Pr(>|t|)
(Intercept) 53.955561 4.586864 11.76306 0.00029883 ***
gdpPercap 0.764883 0.290197 2.63574 0.05783770 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
RMSE: 10.5 Adj. R2: 0.340326Adding another variable
I want to add another fixed effect, so I’m going to add a new column for countries with big vs. small populations.
Running multiple fixed effects
We can add multiple fixed effects with the same syntax as adding more regressors.
OLS estimation, Dep. Var.: lifeExp
Observations: 1,704
Fixed-effects: continent: 5, big_small: 2
Standard-errors: IID
Estimate Std. Error t value Pr(>|t|)
gdpPercap 0.453664 0.023429 19.363 < 2.2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
RMSE: 8.32322 Adj. R2: 0.583092
Within R2: 0.180956Comparing models
fixest has a nice function to print multiple models:
mf mfiid mf2fe
Dependent Var.: lifeExp lifeExp lifeExp
gdpPercap 0.4453*** (0.0235) 0.4453*** (0.0235) 0.4537*** (0.0234)
Fixed-Effects: ------------------ ------------------ ------------------
continent Yes Yes Yes
big_small No No Yes
_______________ __________________ __________________ __________________
S.E. type IID IID IID
Observations 1,704 1,704 1,704
R2 0.57934 0.57934 0.58456
Within R2 0.17456 0.17456 0.18096
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Speed Comparison: fixest and Stata’s reghdfe
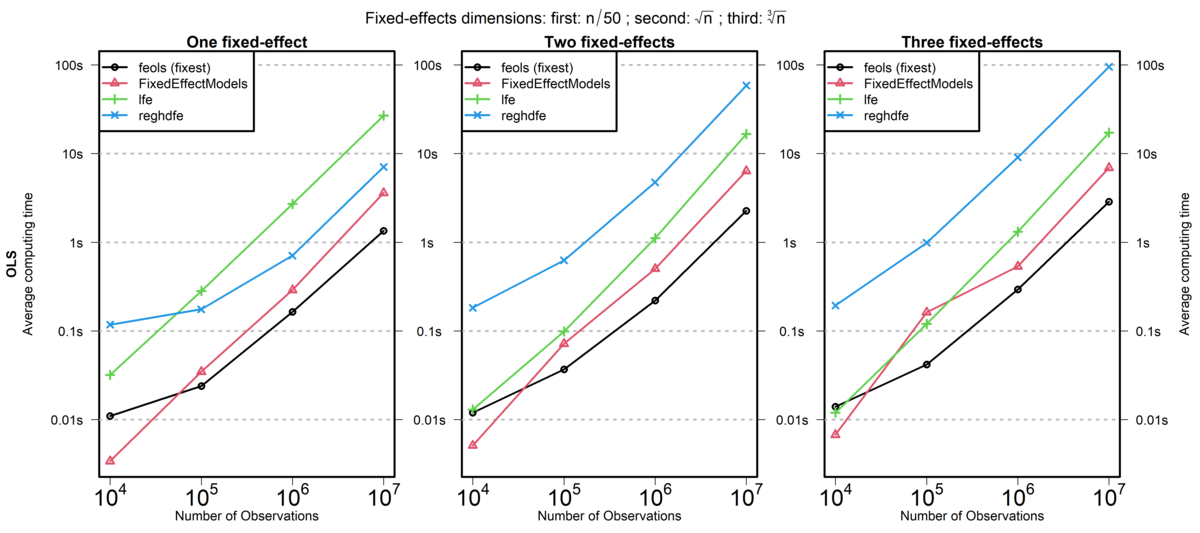
lfe is another R package, reghdfe is a Stata package, and FixedEffectModels is a Julia package.
Source: fixest documentation (comparison run in 2020)
Parallel Processing
Parallel Processing
Sequential processing (default) is when code is run on one “core”.
Parallel Processing
Parallel processing splits operations over multiple “cores”.
A Slow Function
Imagine we had some slow code that we wanted to run multiple times.
Aside: Measuring Code Speed in R
We will use the package tictoc to measure the time it takes to run our function.
If you want more systematic code profiling, you can use the profvis or microbenchmark packages.
Timing our Slow Function
Now we can see how long our slow function takes to run.
Timing our Slow Function
And if we ran it multiple times…
Running in Parallel
If we have a multi-core machine, we can run this slow code in parallel!
Running in Parallel
And this can scale up!
furrr Package
The package we used is called furrr for future and purrr.
future is a package that allows you to run code in parallel.
Documentation:
Setting a Plan
The plan() function sets the parallel processing plan.
sequentialruns code sequentially (the default)multisessionruns code in parallel using multiple R sessionsmulticoreruns code in parallel using multiple cores- Only works on Unix-based systems (Linux, MacOS) and not in Rstudio
Available Cores
You can check how many cores you have available on your machine.
Why Parallel Processing with map()?
Remember that map() is a function that applies another function to each element of a list or vector.
This is always a parallel-izable task.
This is one reason it is good to use map() over a for loop.
- easier to parallelize in the future
Another Example: Bootstrapping
Bootstrapping is a common econometric task that can be easily parallelized.
Let’s generate some data.
Bootstrap Function
To get some standard errors, let’s bootstrap the data.
Bootstrap in Parallel
Or we can run it in parallel!
5.427 sec elapsedPlotting the Results
Plotting the Results
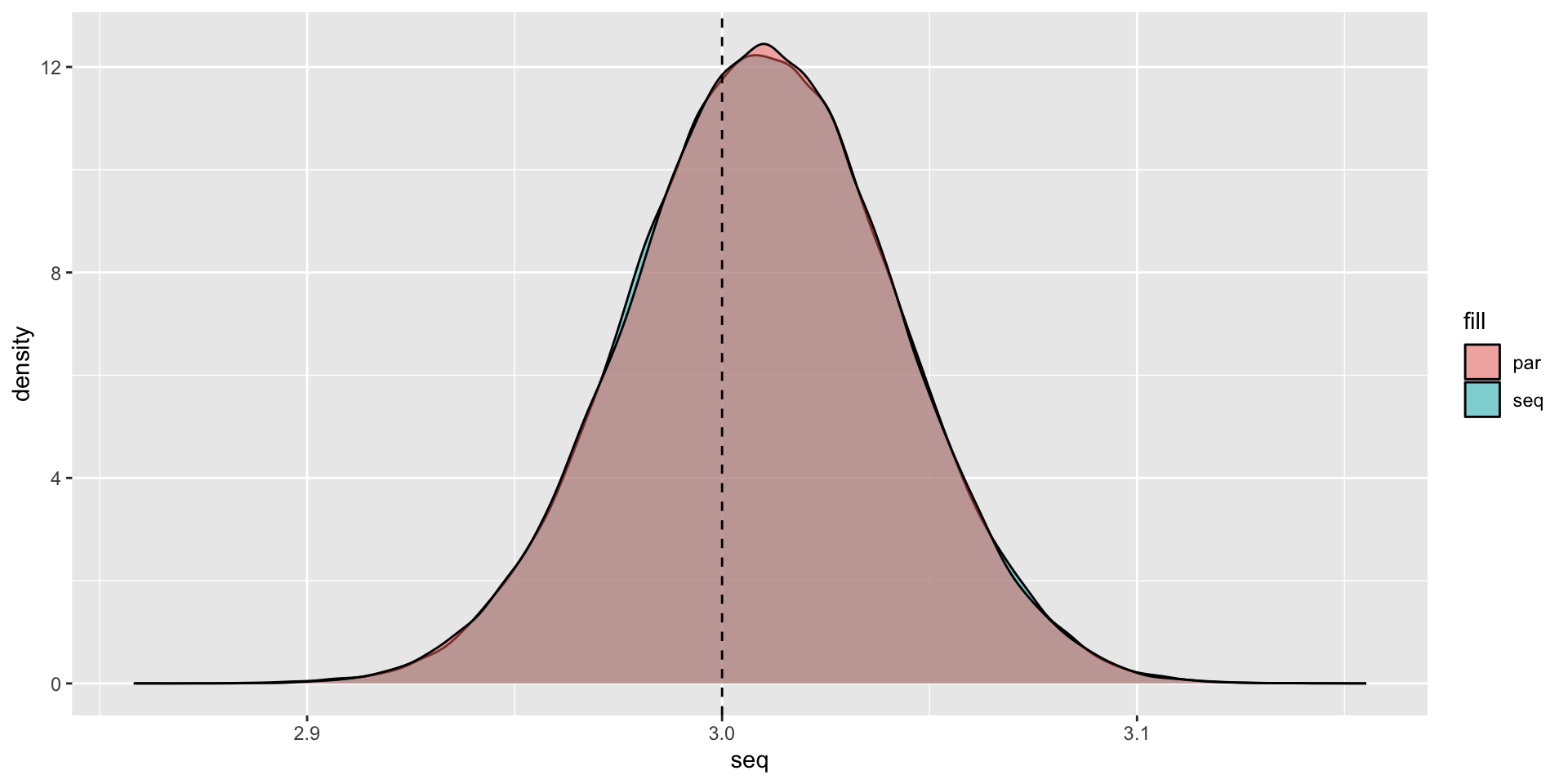
Packages that are already parallelized
A lot of speed performance from packages are from parallel processing.
data.table 1.17.0 using 5 threads (see ?getDTthreads). Latest news: r-datatable.comThis means that adding parallelization on top of data.table operations may not speed things up.
The fixest package also already parallelizes code for multiple fixed effect regressions.
So you have to be careful when paralleizing code that is already optimized.
Back to Functions
When Should You Use a Function?
Rule of Three
- When you duplicate some code three times, you should write it as a function.
This is obviously a rule-of-thumb, but it’s a useful starting point.
You should also consider for your projects,
- will I need to iterate over this step?
- will I need to run robustness checks on this step?
Where Do You Write a Function?
A function has to be declared before you can use it.
So the simplest spot to put them is at the top of your R script.
But this can get overcrowded very quickly.
Sourcing Helper Scripts
A better solution is to store your functions in their own separate scripts.
Then you source() them into the main script, or wherever you need them.
Creating an R Package
Another option is to create an R package for your functions.
This allows you to:
- share your functions with others
- reuse your functions across multiple projects
- add documentation
- add unit tests
You will create a simple test package as part of your homework assignment.
Live Coding Example
Live Coding Example
Live Coding Example - Parallelization on OSCAR
renv::install(c("tibble", "tictoc", "furrr"), prompt = FALSE)
library(tibble)
library(tictoc)
library(furrr)
availableCores()
n = 1e3 ## One thousand observations
set.seed(42)
data <- tibble::tibble(
x = rnorm(n),
y = 4 + 3 * x + rnorm(n)
)
bootstrap <- function(data, n_boot = 1e3) {
boot_data <- data[sample(nrow(data), size = n_boot, replace = TRUE), ]
m <- lm(y ~ x, data = boot_data)
slope <- coef(m)[[2]]
return(slope)
}
plan(multicore)
tic()
slopes_par <- future_map_dbl(1:1e5, ~ bootstrap(data), .options = furrr_options(seed = 42L))
toc()
plan(sequential)
tic()
slopes_par <- future_map_dbl(1:1e5, ~ bootstrap(data), .options = furrr_options(seed = 42L))
toc()